IDENTIFICATION AND BIOCHEMICAL INVESTIGATIONS ON UNKNOWN BACTERIA COLLECTED FROM SEA WATER
HTML Full TextIDENTIFICATION AND BIOCHEMICAL INVESTIGATIONS ON UNKNOWN BACTERIA COLLECTED FROM SEA WATER
P. Rajasulochana * and K. Mukundan
Department of Genetic Engineering, Bharath Institute of Higher Education and Research, Bharath University, Chennai - 600073, Tamil Nadu, India.
ABSTRACT: In marine habitats, a wide diversity of microorganisms such as bacteria, archaea, fungi, and microalgae will exist. Many investigations were reported in the literature on the diversity of marine bacteria and archaea, including their role, usefulness in various domains. It is found from the literature that less work has been reported in the literature related to the role and significance of microbes. Several orders of magnitude between the number of cells will be seen in seawater samples by direct observation and the number of colonies that can grow on agar plates. To prepare subculture, colonies were inoculated in nutrient agar and broth. Bacterial communities are composed of known species which are capable of forming colonies on agar plates but do so with low efficiency. Biochemical investigations were carried out in unknown bacterial samples collected from seawater to know the characteristic of bacteria.
Keywords: Unknown bacteria, Seawater, Biochemical test, Gram’s stain
INTRODUCTION: In most of the cases, when it is required or essential to embark on the process of identifying an organism, it is known that the organism of interest is usually mixed with many other organisms. For instance, to identify the pathogen which causes a disease in a patient, there are many different species of bacteria present in the stool or tissue sample that are to be obtained from the patient. If the organism is not in pure form, the first step would be to separate it from all the others. This can be done in two stages.
The first stage, which is called primary isolation, is to identify the appropriate media, time, and temperature of incubation to ensure that the organism of interest has a good opportunity of being isolated. Knowledge of the specimen source and a microscopic description of the mixture are very much required in choosing the proper media and growth conditions. For example, if a stool specimen of a patient is being examined for a possible pathogen, a Mac Conkey plate must be included for use in the primary isolation.
Since pathogens in stool samples are primarily Gram-negative rods; the MacConkey would increase the likelihood of their isolation because it selects for this group of organisms. The second stage is the secondary isolation where the subculturing of each colony type from the primary isolation onto non-inhibitory media is accomplished. Sometimes multiple sub culturing’s are necessary to obtain a pure secondary culture. Upon isolation of a pure culture, each organism is described in detail according to cellular and colonial morphologies, optimum growth characteristics, and Gram stain reactions. Growth on a plate can distort the characteristic three-dimensional cell arrangement, whereas a broth allows the direction of a cell division to occur naturally in different planes. Thus, cellular descriptions of size, shape, and arrangement must be made from a broth culture. Also, an accurate measure of cell length and width is very important because it may be the only definitive characteristic available to distinguish one bacterium from another. Thus to have a basic framework to start from, we need information on the shape (rod, coccus, spirillum) and Gram stain reaction (positive, negative) of the species of interest.
In the lab, identification of unknown bacteria is divided into 3 projects; EU, OU, and PCR-ID projects. For EU (identification of a bacterial species belonging to the Enterobacteriaceae family) and OU (identification of a non-Enterobacteriaceae bacterial species), we use the conventional microbiological diagnostic techniques to identify the species. For the PCR-ID project, we will be using modern molecular biology techniques whereby a specific part of the genomic DNA is amplified, and its sequence of bases is determined and compared to those of different bacterial species to find a match. You will be dealing with each one of these unknown projects in their proper time and place, but for each one, we require that you write a paper on the subject. It is possible that your instructor may decide that you perform the PCR-ID on the same sample that you obtained as your OU or EU, in which case only two reports will suffice.
MATERIALS AND METHODS:
Sample Collection: Seawater samples were collected in a sterilized container from the sea at a depth of 1m and 6 km away from the seashore at Thiruvanmiyur in Tamil Nadu, India.
Nutrient Agar Preparation:
- Suspend 28 g of nutrient agar powder in 1 liter of distilled water.
- Heat this mixture while stirring to fully dissolve all components.
- Autoclave the dissolved mixture at 121 degrees Celsius for 15 min.
- Once the nutrient agar has been autoclaved, allow it to cool but not solidify.
- Pour nutrient agar into each plate and leave plates on the sterile surface until the agar has solidified.
- Replace the lid of each Petri dish and store the plates in a refrigerator.
FIG. 1: NUTRIENT AGAR PREPARATION
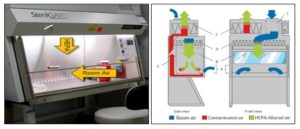
Laminar Air Procedure: A laminar flow cabinet or laminar flow closet or tissue culture hood is a carefully enclosed bench designed to prevent contamination of semiconductor wafers, biological samples, or any particle sensitive materials. Air is drawn through a HEPA filter and blown in a very smooth, laminar flow towards the user. Due to the direction of air flow, the sample is protected from the user, but the user is not protected from the sample. The cabinet is usually made of stainless steel with no gaps or joints where spores might collect. Such hoods exist in both horizontal and vertical configurations, and there are many different types of cabinets with a variety of airflow patterns and acceptable uses. Laminar flow cabinets may have a UV-C germicidal lamp to sterilize the interior and contents before usage to prevent contamination of experiment. Germicidal lamps are usually kept on for 15 min to sterilize the interior, and no contact is to be made with a laminar flow hood during this time. During this time, investigators normally prepare other materials to maximize efficiency. It is important to switch this light off during use, to limit exposure to skin and eyes as stray ultraviolet light emissions can cause cancer and cataracts.
FIG. 2: LAMINAR AIR
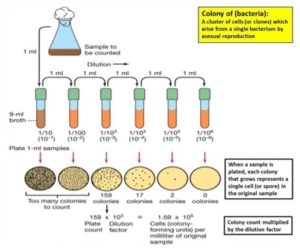
Serial Dilution Technique: A serial dilution is a series of sequential dilutions used to reduce a dense culture of cells to a more usable concentration. Each dilution will reduce the concentration of bacteria by a specific amount. So, by calculating the total dilution over the entire series, it is possible to know many bacteria you started with. To start, we need 10 milliliters (10 ml) of your original bacterial culture (labeled OBC). Before we start diluting, we need to prepare several dilution blanks, which are tubes containing your diluting liquid in exact quantities. Your liquid could be growth media, saline, sterile water, or any other appropriate liquid. For this example, we need 5 dilution blanks, numbered 1-5. In each tube, we need exactly 9 ml of liquid media. The reason we need 9 ml will become apparent soon. The first step is to gently shake or swirl the tube.
This will ensure that your cells are evenly distributed in the tube. If your cells settle to the bottom, and you remove liquid without swirling, you run the risk of not getting enough cells, invalidating your final count. Remember to always swirl the tube before removing liquid. Once swirled, carefully transfer exactly 1 ml from your OBC Tube to Tube 1. Now, you should have 10 ml of liquid in Tube 1. Exactly one-tenth of your cells are now in a new tube with a final volume of 10 ml. You just performed a 1 in 10 dilutions, or it could be written 1/10. 1 is the volume you transferred, and 10 is the final volume of the tube after the transfer.
FIG. 3: SERIAL DILUTION TECHNIQUE
Estimation of Total Salt Concentration and pH of the Water Samples: About 10 ml of each sample were taken in a two separate clean, and dry test tube whose weight is determined previously and they were allowed to evaporate till there is no trace of water droplets inside the test tubes. The weight of the test tubes with the salts was measured, and the difference between the weight of the test tube with salt and the empty test tube gives the weight of the salts present in the water samples. The pH of the water samples was determined using pH meter, and values were recorded.
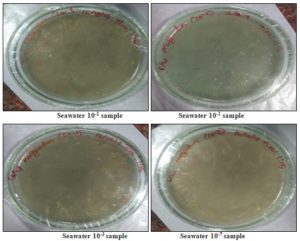
Isolation of Sea Water Samples: The samples serially diluted in the range of 10-1, 10-2, 10-3, 10-4, 10-5, 10-6, and 10-7 in a series of the test tube. The cultured in the nutrient medium with an increased concentration of NaCl.
The pH of the medium was adjusted to 7-7.4. The samples were inoculated under the aseptic condition in the Petri plates containing nutrient agar medium using spread plate technique and are incubated at 37 °C for 48 h 1.
Bacterial Growth:
Seawater Mother Culture 4 Petri plates:
Seawater Sap Culture Six Petri plates:
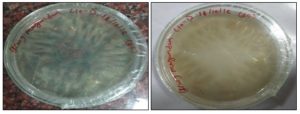
FIG. 4: SAP CULTURE 10-1 PETRI PLATES ddb Scb
FIG. 5: SAP CULTURE 10-2 PETRI PLATES
FIG. 6: SAP CULTURE 10-3 PETRI PLATES
Observation of Bacterial Colony Color: (x) White colony, (x) Dirty white colony, (x) Spongy white colony, (x) Milky white colony, (x) Yellow pigmented colony.
Gram Staining: The morphological studies like colony appearance, cell shape, and size on the obtained cultures were carried out using Gram staining method.
The stained slides were observed for the morphological characteristics of the culture under the compound microscope.
Biochemical Characterization: Different biochemical tests were performed to characterize and to confirm the isolated species as Halobacterium, such as catalase test, indole production test, methyl red, and vogas proskauer (MR – VP) test, citrate utilization test and acid production test.
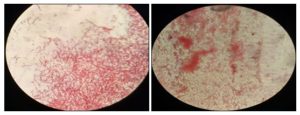
FIG. 7: ADDED TO SLIDE 1 AND SLIDE 2 PHOTOS. IT IS GRAM-NEGATIVE BACTERIA
RESULTS AND DISCUSSION: Identification of unknown bacteria in seawater samples have “Halobacterium” pep. It is gram-negative bacteria find out for Gram staining method and microscope.
Isolation of Halobacterium: Numerous colonies like yellow pigmented, red pigmented, white spongy, white radial, dirty white colonies were observed from the water samples. The isolated Halobacterium sp. from the seawater samples was found as radial white spongy gram-negative bacillus colonies. Ninety-five extremely halophilic species have been isolated from distinct saline regions of Turkey. Likewise, a novel species of saltwater bacterial, namely Halobacterium jilantaiense was isolated.
Morphological Characteristics of Halobacterium: The isolated Halobacterium strains appeared as rod-like Bacillus, gram-negative organism. They found as radial white spongy colonies. The results of the morphological study of the obtained colonies were indicated in Table 1. All the colonies isolated both from the seawater and paddy field water were gram-negative bacilli. They are motile and aerobic, requires oxygen for their metabolism.
Determination of Type of Respiration in Halobacterium sp.: It was observed that the colonial growth was absent in the test tubes sealed with the waxes due to the aerobic nature of the colony which requires oxygen for its respiration. Both the aerobic and anaerobic colonies were isolated from water samples. The results were indicated.
Biochemical Characteristics of Halobacterium sp.: The isolated pure cultures are subjected to various biochemical characterization, and the results were indicated.
The bacterial colonies isolated from the seawater samples were showed positive results almost for all the biochemical tests except a few carried out in the present study. The results of the morphological and biochemical characteristic studies have confirmed the Halobacterium sp.
TABLE 1: MORPHOLOGICAL CHARACTERISTICS OF HALOBACTERIUM
| S. no. | Sample | Colony Appearance | Cell shape | Gram staining |
| 1 | 10-1 (i) | White colony | Bacilli | Gram-negative |
| 2 | 10-1 (ii) | Dirty white colony | Bacilli | Gram-negative |
| 3 | 10-2 (i) | Spongy white colony | Bacilli | Gram-negative |
| 4 | 10-2 (ii) | Milky white colony | Bacilli | Gram-negative |
| 5 | 10-3 (i) | Yellow pigmented colony | Bacilli posses terminal spores | Gram-negative |
| 6 | 10-3 (ii) | White colony | Bacilli | Gram-negative |
TABLE 2: DETERMINATION OF TYPE OF RESPIRATION IN HALOBACTERIUM SP.
| S. no. | Sample | Colony Appearance | Respiration Type |
| 1 | 10-1 (i) | White colony | Aerobes |
| 2 | 10-1 (ii) | Dirty white colony | Aerobes |
| 3 | 10-2 (i) | Spongy white colony | Aerobes |
| 4 | 10-2 (ii) | Milky white colony | Facultative anaerobes |
| 5 | 10-3 (i) | Yellow pigmented colony | Aerobes |
| 6 | 10-3 (ii) | White colony | Facultative anaerobes |
TABLE 3: BIOCHEMICAL CHARACTERISTICS OF HALOBACTERIUM SP.
| Name of Biochemical test | 10-1 (i) | 10-1 (ii) | 10-2 (i) | 10-2 (ii) | 10-3 (i) | 10-3 (ii) |
| Catalase test | + | + | + | + | + | + |
| Indole production test | + | + | + | + | + | + |
| Methyl red test | + | + | + | - | + | + |
| Vogas Proskauer test | - | - | - | + | - | - |
| Citrate utilization test | + | + | + | + | + | + |
| Acid production test | + | + | + | + | + | + |
CONCLUSION: Numerous colonies such as (i) yellow pigmented, (ii) red pigmented, (iii) white spongy, (iv) white radial, (v) dirty white colonies were observed from the water samples. Identification of unknown bacteria in seawater samples is found to have “Halobacterium” pep. It is gram-negative bacteria, the output realized through the Gram staining method and microscope. The isolated Halobacterium strains visually appeared as rod-like Bacillus, gram-negative organism. They found in the form of radial white spongy colonies. It is observed that the colonial growth was absent in the test tubes sealed with the waxes due to the aerobic nature of the colony, which essentially requires oxygen for its respiration. Both the aerobic and anaerobic colonies are isolated from water samples. The isolated pure cultures are subjected to various biochemical characterization, and the results are consolidated. The bacterial colonies isolated from the seawater samples are exhibited positive results almost for all the biochemical tests except a few carried out in the present study.
SUMMARY: A wide diversity of microorganisms such as bacteria, archaea, fungi, and microalgae are found in marine habitats. Although there has been much progress about the diversity of marine bacteria and archaea, there exists less literature about the “role” of these microbes. Several orders of magnitude between the number of cells that can be seen in seawater samples by direct observation and the number of colonies that grows on agar plates. Colonies were inoculated in nutrient agar and broth for subculture.
Bacterial communities are composed of known species which are capable of forming colonies on agar plates but do so with low efficiency. Biochemical tests were carried out on unknown bacterial samples collected from seawater. Identification of unknown bacteria in seawater samples have “Halobacterium” pep. It is gram-negative bacteria found for Gram staining method and microscope.
ACKNOWLEDGEMENT: Nil
CONFLICT OF INTEREST: Nil
REFERENCES:
- Seelay HWJ and Vandermark PJ: Microbes in Action. A laboratory Manual of Microbiology, W. H. Freeman & Co., 1981: 135.
- Yang Y, Cui HL, Zhou PJ and Liu SJ: Halobacterium jilantaiense nov., a halophilic archaeon isolated from a saline lake in Inner Mongolia, China. Int J Syst Evol Microbiol 2006; 56 (pt10): 2353-5.
- Dym O, Mevarech M and Sussman JL: Structural features that stabilize halphilic malate dehydrogenase from an archaebacterium. Science 1995; 267: 1344-46.
- Galinski EA: Copatible solutes of halophilic eubacteria: molecular principles, water - soluteinteractions, stress protection. Experientia 1993; 49: 487-96.
- Gibbons NE: Isolation, growth and requirements of Halophilic bacteria. In F.N. Norris and D.W. Ribbons (ed.), Methods in microbiology, Academic press Inc., Newyork, Vol. 3B, 1969. 169-83.
- GonZalez C, Gutierrez C and Ramirez C: Halobacterium vallismortis Nov. an amylolytic and carbohydrate metabolizing, extremely halophilic bacterium. Can J Microbiol 1978, 24: 710-15.
- Ha DJ, Joo WA, Han GY and Kim CW: Proteome analysis of Halobacterium salinarum and characterization of proteins related to the degradation of isopropyl alcohol. Biochem Biophys Acta 2007; 1774(1): 44-50.
How to cite this article:
Rajasulochana P and Mukundan K: Identification and biochemical investigations on unknown bacteria collected from sea water. Int J Life Sci & Rev 2019; 5(3): 42-48. doi: 10.13040/IJPSR.0975-8232.IJLSR.5(3).42-48.
All © 2015 are reserved by International Journal of Life Sciences and Review. This Journal licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License.
Article Information
2
42-48
718
1281
English
IJLSR
P. Rajasulochana * and K. Mukundan
Department of Genetic Engineering, Bharath Institute of Higher Education and Research, Bharath University, Chennai, Tamil Nadu, India.
prsnellore@gmail.com
01 March 2019
15 March 2019
22 March 2019
10.13040/IJPSR.0975-8232.IJLSR.5(3).42-48
31 March 2019